Journal of Fisheries Research ›› 2023, Vol. 45 ›› Issue (4): 331-340.DOI: 10.14012/j.cnki.fjsc.2023.04.002
• Research Paper • Previous Articles Next Articles
LI Zhaonan( ), LI Changzhong, BAO Changhong, HE Caixia, JIN Wenjie, CHEN Yanxia(
), LI Changzhong, BAO Changhong, HE Caixia, JIN Wenjie, CHEN Yanxia( )
)
Received:2022-12-05
Online:2023-08-25
Published:2023-08-09
李昭楠( ), 李长忠, 保长虹, 贺彩霞, 金文杰, 陈艳霞(
), 李长忠, 保长虹, 贺彩霞, 金文杰, 陈艳霞( )
)
通讯作者:
陈艳霞(1987—),女,博士,讲师,主要从事高原动物保护与利用相关研究。E-mail:chenyanxia9568@163.com
作者简介:李昭楠(1996—),女,硕士,主要从事动物生态学相关研究。E-mail:lizhaonan123456@163.com
基金资助:CLC Number:
LI Zhaonan, LI Changzhong, BAO Changhong, HE Caixia, JIN Wenjie, CHEN Yanxia. Molecular identification of Oncorhynchus mykiss based on DNA barcode technology[J]. Journal of Fisheries Research, 2023, 45(4): 331-340.
李昭楠, 李长忠, 保长虹, 贺彩霞, 金文杰, 陈艳霞. 基于DNA条形码技术的虹鳟分子鉴定[J]. 渔业研究, 2023, 45(4): 331-340.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.hyyysci.com/EN/10.14012/j.cnki.fjsc.2023.04.002
| 物种 Species | 登录号 Accession number | 长度/bp Length |
|---|---|---|
| 帝王鲑 Oncorhynchus tshawytscha | KX958414.1 | 1 204 |
| 金腹大麻哈鱼 Oncorhynchus chrysogaster | JX960908.1 | 1 247 |
| 美洲红点鲑 Salvelinus fontinalis | HQ167687.1 | 1 551 |
| 钝吻鲑 Salmo obtusirostris | JX960948.1 | 1 262 |
| 多瑙哲罗鱼 Hucho hucho | JX960904.1 | 1 247 |
| 细鳞鲑 Brachymystax lenok | JX227988.1 | 1 581 |
| 突唇白鲑 Coregonus lavaretus | JX960890.1 | 1 246 |
| 北鲑 Stenodus leucichthys | KT630716.1 | 1 201 |
| 普伦白鲑 Coregonus pollan | JX960898.1 | 1 262 |
| 茴鱼 Thymallus thymallus | JX960975.1 | 1 262 |
Tab.1 COⅠgene information of Salmonidae
| 物种 Species | 登录号 Accession number | 长度/bp Length |
|---|---|---|
| 帝王鲑 Oncorhynchus tshawytscha | KX958414.1 | 1 204 |
| 金腹大麻哈鱼 Oncorhynchus chrysogaster | JX960908.1 | 1 247 |
| 美洲红点鲑 Salvelinus fontinalis | HQ167687.1 | 1 551 |
| 钝吻鲑 Salmo obtusirostris | JX960948.1 | 1 262 |
| 多瑙哲罗鱼 Hucho hucho | JX960904.1 | 1 247 |
| 细鳞鲑 Brachymystax lenok | JX227988.1 | 1 581 |
| 突唇白鲑 Coregonus lavaretus | JX960890.1 | 1 246 |
| 北鲑 Stenodus leucichthys | KT630716.1 | 1 201 |
| 普伦白鲑 Coregonus pollan | JX960898.1 | 1 262 |
| 茴鱼 Thymallus thymallus | JX960975.1 | 1 262 |
| 引物名称 Name of primers | 引物序列 Sequence of primers | 产物长度/bp Length of product | |
|---|---|---|---|
| COⅠ-1(DNA barcode) | COⅠ-F24 | 5’-GGGATGACCAAATCTATAACGTGA-3’ | 681 |
| COⅠ-R24 | 5’-CTCAGACCATTCCTATATACCCGAAG-3’ | ||
| COⅠ-F25 | 5’-GATTAATTCCCCTAATAATCGGAGC-3’ | 678 | |
| COⅠ-R25 | 5’-ATGTAAAGTAAGCACGAGTGTCCA-3’ | ||
| COⅠ-2(Mini barcode) | TEYI-0F | 5’-AACCTCCAGCCATCTCTCAG-3’ | 153 |
| TEYI-0R | 5’-CCGGGTCAAAGAAAGTGGTG-3’ | ||
| TEYI-1F | 5’-CCGCCCTGAGTCTACTGATT-3’ | 230 | |
| TEYI-1R | 5’-TGGAGGAAGGAGTCAGAAGC-3’ | ||
Tab.2 PCR primer information
| 引物名称 Name of primers | 引物序列 Sequence of primers | 产物长度/bp Length of product | |
|---|---|---|---|
| COⅠ-1(DNA barcode) | COⅠ-F24 | 5’-GGGATGACCAAATCTATAACGTGA-3’ | 681 |
| COⅠ-R24 | 5’-CTCAGACCATTCCTATATACCCGAAG-3’ | ||
| COⅠ-F25 | 5’-GATTAATTCCCCTAATAATCGGAGC-3’ | 678 | |
| COⅠ-R25 | 5’-ATGTAAAGTAAGCACGAGTGTCCA-3’ | ||
| COⅠ-2(Mini barcode) | TEYI-0F | 5’-AACCTCCAGCCATCTCTCAG-3’ | 153 |
| TEYI-0R | 5’-CCGGGTCAAAGAAAGTGGTG-3’ | ||
| TEYI-1F | 5’-CCGCCCTGAGTCTACTGATT-3’ | 230 | |
| TEYI-1R | 5’-TGGAGGAAGGAGTCAGAAGC-3’ | ||
| 种名 Species | 登录号 Accession number | 长度/bp Length | |
|---|---|---|---|
| 帝王鲑 Oncorhynchus tshawytscha | FJ999371.1 | 652 | |
| 银鲑 Oncorhynchus kisutch | MG951604.1 | 652 | |
| 虹鳟 Oncorhynchus mykiss | MN850431.1 | 653 | |
| 大麻哈鱼 Oncorhynchus keta | MN850432.1 | 653 | |
| 大西洋鲑 Salmo salar | MN850430.1 | 651 | |
| 花羔红点鲑 Salvelinus malma | EU522414.1 | 652 | |
| 北极红点鲑 Salvelinus alpinus | KJ128606.1 | 648 | |
| 远东红点鲑 Salvelinus leucomaenis | MF503661.1 | 697 | |
| 多瑙哲罗鱼 Hucho hucho | KJ553640.1 | 652 | |
| 哲罗鱼 Hucho taimen | MG951558.1 | 652 | |
| 细鳞鲑 Brachymystax lenok | JX261990.1 | 633 | |
| 远东哲罗鱼 Parahucho perryi | HQ693237.1 | 655 | |
| 秋白鲑 Coregonus autumnalis | EU202649.1 | 650 | |
| 高白鲑 Coregonus peled | MF632325.1 | 637 | |
| 欧白鲑 Coregonus albula | KX457959.1 | 704 | |
| 茴鱼 Thymallus thymallus | HQ961016.1 | 652 | |
| 黑龙江茴鱼 Thymallus grubii | MG951577.1 | 652 | |
Tab.3 COⅠ gene information of 17 Salmonidae species
| 种名 Species | 登录号 Accession number | 长度/bp Length | |
|---|---|---|---|
| 帝王鲑 Oncorhynchus tshawytscha | FJ999371.1 | 652 | |
| 银鲑 Oncorhynchus kisutch | MG951604.1 | 652 | |
| 虹鳟 Oncorhynchus mykiss | MN850431.1 | 653 | |
| 大麻哈鱼 Oncorhynchus keta | MN850432.1 | 653 | |
| 大西洋鲑 Salmo salar | MN850430.1 | 651 | |
| 花羔红点鲑 Salvelinus malma | EU522414.1 | 652 | |
| 北极红点鲑 Salvelinus alpinus | KJ128606.1 | 648 | |
| 远东红点鲑 Salvelinus leucomaenis | MF503661.1 | 697 | |
| 多瑙哲罗鱼 Hucho hucho | KJ553640.1 | 652 | |
| 哲罗鱼 Hucho taimen | MG951558.1 | 652 | |
| 细鳞鲑 Brachymystax lenok | JX261990.1 | 633 | |
| 远东哲罗鱼 Parahucho perryi | HQ693237.1 | 655 | |
| 秋白鲑 Coregonus autumnalis | EU202649.1 | 650 | |
| 高白鲑 Coregonus peled | MF632325.1 | 637 | |
| 欧白鲑 Coregonus albula | KX457959.1 | 704 | |
| 茴鱼 Thymallus thymallus | HQ961016.1 | 652 | |
| 黑龙江茴鱼 Thymallus grubii | MG951577.1 | 652 | |
| 物种 Species | 浓度/(ng/μL) Concentration | OD(260/280) | 260/230 | 260 |
|---|---|---|---|---|
| 三倍体虹鳟Triploid rainbow trout | 164.90 | 2.032 | 1.343 | 3.300 |
| 金鳟Golden trout | 197.65 | 2.141 | 1.633 | 3.964 |
| 二倍体虹鳟 Diploid rainbow trout | 119.85 | 2.001 | 1.097 | 2.366 |
| 大西洋鲑Salmo salar | 170.00 | 1.988 | 1.943 | 3.406 |
| 大麻哈鱼Oncorhynchus keta | 198.40 | 2.084 | 1.859 | 3.957 |
| 高白鲑Coregonus peled | 126.05 | 1.982 | 2.030 | 2.507 |
Tab.4 Test results of DNA quality of various samples
| 物种 Species | 浓度/(ng/μL) Concentration | OD(260/280) | 260/230 | 260 |
|---|---|---|---|---|
| 三倍体虹鳟Triploid rainbow trout | 164.90 | 2.032 | 1.343 | 3.300 |
| 金鳟Golden trout | 197.65 | 2.141 | 1.633 | 3.964 |
| 二倍体虹鳟 Diploid rainbow trout | 119.85 | 2.001 | 1.097 | 2.366 |
| 大西洋鲑Salmo salar | 170.00 | 1.988 | 1.943 | 3.406 |
| 大麻哈鱼Oncorhynchus keta | 198.40 | 2.084 | 1.859 | 3.957 |
| 高白鲑Coregonus peled | 126.05 | 1.982 | 2.030 | 2.507 |
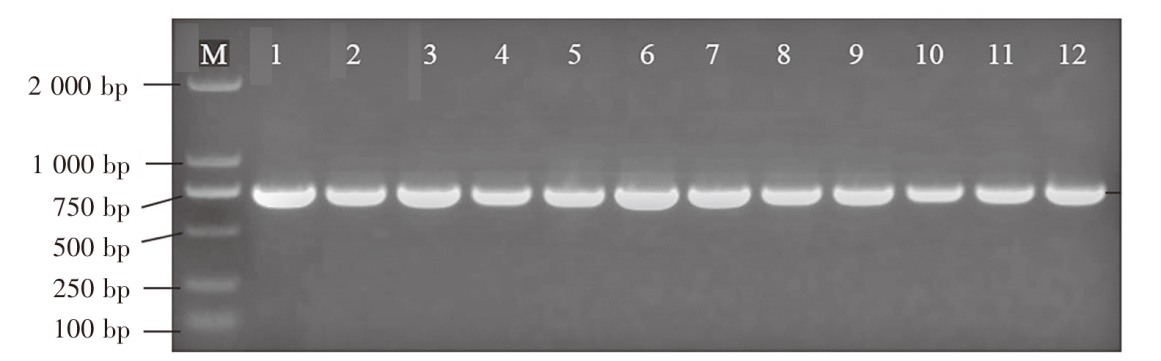
Fig.1 DNA barcode PCR results Notes: M indicated DL 2000 molecular quality standard; 1-6 used COⅠ-F24 and COⅠ-R24 primers; 7-12 used COⅠ-F25 and COⅠ-R25 primers; 1 and 7 were triploid rainbow trout; 2 and 8 were golden trout; 3 and 9 were diploid rainbow trout; 4 and 10 were S.salar; 5 and 11 were O.keta; 6 and 12 were C.peled.
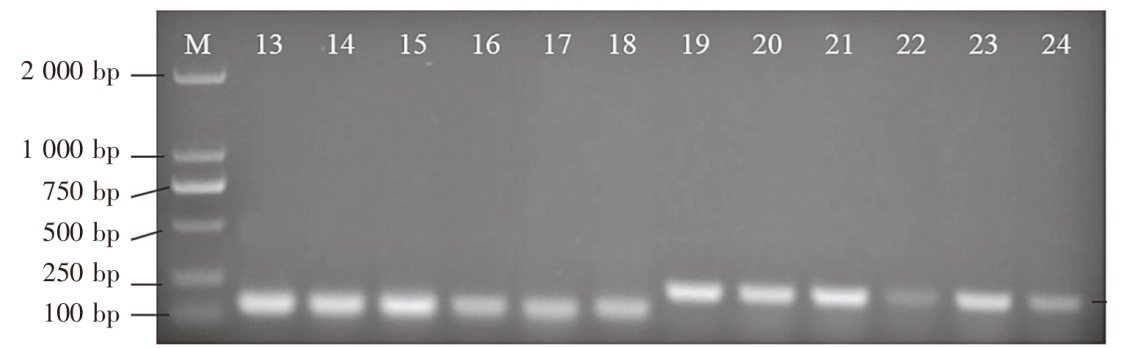
Fig.2 Mini barcode PCR results Notes: M indicated DL 2000 molecular quality standard; 13-18 used TEYI-0F and TEYI-0R primers; 19-24 used TEYI-1F and TEYI-1R primers; 13 and 19 were triploid rainbow trout; 14 and 20 were golden trout; 15 and 21 were diploid rainbow trout; 16 and 22 were S.salar; 17 and 23 were O.keta; 18 and 24 were C.peled.
| 标签名称 Label name | DNA barcode鉴定结果 (序列相似度/%) DNA barcode identification results (sequence similarity) | Mini barcode鉴定结果 (序列相似度/%) Mini barcode identification results (sequence similarity) | 结合两个片段鉴定结果 Combining the identification results of the two fragment |
|---|---|---|---|
| 三倍体虹鳟 Triploid rainbow trout | 虹鳟(100) | 虹鳟(98.69) | 虹鳟 |
| 金鳟Golden trout | 虹鳟(100) | 虹鳟(98.69) | 虹鳟 |
| 二倍体虹鳟 Diploid rainbow trout | 虹鳟(100) | 虹鳟(98.69) | 虹鳟 |
| 大西洋鲑Salmo salar | 大西洋鲑(100) | 大西洋鲑(100) | 大西洋鲑 |
| 大麻哈鱼 Oncorhynchus keta | 大麻哈鱼(100) | 大麻哈鱼(97.06) | 大麻哈鱼 |
| 高白鲑 Coregonus peled | 欧白鲑(100) | 欧白鲑(98.52) | 欧白鲑 |
Tab.5 Identification results of NCBI database of 6 species of Salmonidae by DNA barcode technology
| 标签名称 Label name | DNA barcode鉴定结果 (序列相似度/%) DNA barcode identification results (sequence similarity) | Mini barcode鉴定结果 (序列相似度/%) Mini barcode identification results (sequence similarity) | 结合两个片段鉴定结果 Combining the identification results of the two fragment |
|---|---|---|---|
| 三倍体虹鳟 Triploid rainbow trout | 虹鳟(100) | 虹鳟(98.69) | 虹鳟 |
| 金鳟Golden trout | 虹鳟(100) | 虹鳟(98.69) | 虹鳟 |
| 二倍体虹鳟 Diploid rainbow trout | 虹鳟(100) | 虹鳟(98.69) | 虹鳟 |
| 大西洋鲑Salmo salar | 大西洋鲑(100) | 大西洋鲑(100) | 大西洋鲑 |
| 大麻哈鱼 Oncorhynchus keta | 大麻哈鱼(100) | 大麻哈鱼(97.06) | 大麻哈鱼 |
| 高白鲑 Coregonus peled | 欧白鲑(100) | 欧白鲑(98.52) | 欧白鲑 |
| [1] |
刘小鹏, 马存霞, 魏丽, 等. 黄河上游地区减贫转向与高质量发展[J]. 资源科学, 2020, 42(1):197-205.
DOI |
| [2] | 简生龙. 基于生态优先前提下推动青海鲑鳟鱼养殖绿色发展的探讨[J]. 青海农林科技, 2019(3):78-81. |
| [3] | 简生龙, 关弘弢, 李柯懋, 等. 青海沿黄鲑鳟鱼网箱养殖水体环境监测研究[J]. 中国水产, 2020(5):53-58. |
| [4] |
Berthelot C, Brunet F, Chalopin D, et al. The rainbow trout genome provides novel insights into evolution after whole-genome duplication in vertebrates[J]. Nature Communication, 2014, 5: 3657.
DOI |
| [5] | Habte-Tsion H, Ren M, Liu B, et al. Threonine modulates immune response, antioxidant status and gene expressions of antioxidant enzymes and antioxidant-immune-cytokine-related signaling molecules in juvenile blunt snout bream (Megalobrama amblycephala)[J]. Fish & Shellfish Immunology, 2016, 51: 189-199. |
| [6] |
Meiler K A, Kumar V. Organic and inorganic zinc in the diet of a commercial strain of diploid and triploid rainbow trout (Oncorhynchus mykiss): effects on performance and mineral retention[J]. Aquaculture, 2021, 545: 737126.
DOI URL |
| [7] |
Gonçalves J F M, Hinzmann M, Machado J, et al. Oxidative effect of L-carnitine on energy metabolism in diploid and triploid rainbow trout (Oncorhynchus mykiss): impact on metabolites[J]. International Aquatic Research, 2018, 10(2): 133-143.
DOI |
| [8] |
Huang T, Gu W, Liu E, et al. Comprehensive analysis of miRNA-mRNA/lncRNA during gonadal development of triploid female rainbow trout (Oncorhynchus mykiss)[J]. Genomics, 2021, 113(6): 3533-3543.
DOI PMID |
| [9] |
Xu G, Huang T, Gu W, et al. Effects of letrozole and 17α-methyltestosterone on gonadal development in all-female triploid rainbow trout (Oncorhynchus mykiss)[J]. Aquaculture Research, 2021, 52(6): 2460-2469.
DOI URL |
| [10] |
Meiler K A, Cleveland B, Radler L, et al. Oxidative stress-related gene expression in diploid and triploid rainbow trout (Oncorhynchus mykiss) fed diets with organic and inorganic zinc[J]. Aquaculture, 2021, 533: 736149.
DOI URL |
| [11] |
Weber G M, Ma H, Birkett J, et al. Effects of feeding level and sexual maturation on expression of genes regulating growth mechanisms in rainbow trout (Oncorhynchus mykiss)[J]. Aquaculture, 2022, 551: 737917.
DOI URL |
| [12] |
Carvalho D C, Palhares R M, Drummond M G, et al. DNA barcoding identification of commercialized seafood in South Brazil: a governmental regulatory forensic program[J]. Food Control, 2015, 50: 784-788.
DOI URL |
| [13] |
Hwang I K, Lee H Y, Kim M, et al. Development of real-time PCR assay for genetic identification of the mottled skate, Beringraja pulchra[J]. Forensic Science International, 2015, 255: 80-84.
DOI PMID |
| [14] | 时圣明, 潘明佳, 王洁, 等. 分子鉴定技术在中药中的应用[J]. 中草药, 2016, 47(17):3121-3126. |
| [15] |
Handy S M, Deeds J R, Ivanova N V, et al. A single-laboratory validated method for the generation of DNA barcodes for the identification of fish for regulatory compliance[J]. Journal of AOAC International, 2011, 94(1): 201-210.
PMID |
| [16] |
Rhee J S, Seo J S, Raisuddin S, et al. Gonadotropin-releasing hormone receptor (GnRHR) gene expression is differently modulated in gender types of the hermaphroditic fish Kryptolebias marmoratus by endocrine disrupting chemicals[J]. Comparative Biochemistry and Physiology C Toxicology Pharmacology, 2008, 147(3): 357-365.
DOI URL |
| [17] |
Ash K T, Drake K M, Gibbs W S, et al. Genomic diversity of type B3 bacteriophages of Caulobacter crescentus[J]. Current Microbiology, 2017, 74(7): 779-786.
DOI |
| [18] |
Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets[J]. Molecular Biology and Evolution, 2016, 33(7): 1870-1874.
DOI PMID |
| [19] |
Jaffe A, Amsel N, Aizenbud Y, et al. Spectral neighbor joining for reconstruction of latent tree models[J]. SIAM Journal on Mathematics of Data Science, 2021, 3(1):113-141.
DOI PMID |
| [20] |
Dudgeon C L, Blower D C, Broderick D, et al. A review of the application of molecular genetics for fisheries management and conservation of sharks and rays[J]. Journal of Fish Biology, 2012, 80(5): 1789-1843.
DOI PMID |
| [21] |
Teletchea F. Molecular identification methods of fish species: reassessment and possible applications[J]. Reviews in Fish Biology and Fisheries, 2009, 19(3): 265-293.
DOI URL |
| [22] | 衣洁菡, 王桐, 郭德慧, 等. 利用SNP分子标记对阿根廷滑柔鱼和科氏滑柔鱼的分子鉴定[J]. 食品科技, 2018, 43(10):311-316. |
| [23] |
Kress W J, García-Robledo C, Uriarte M, et al. DNA barcodes for ecology,evolution,and conservation[J]. Trends in Ecology & Evolution, 2015, 30(1):25-35.
DOI URL |
| [24] |
Hulley E N, Taylor N D J, Zarnke A M, et al. DNA barcoding vs. morphological identification of larval fish and embryos in Lake Huron: advantages to a molecular approach[J]. Journal of Great Lakes Research, 2018, 44(5): 1110-1116.
DOI URL |
| [25] |
Nedunoori A, Turanov S V, Kartavtsev Y P. Fish product mislabeling identified in the Russian far east using DNA barcoding[J]. Gene Reports, 2017, 8: 144-149.
DOI URL |
| [26] |
Cline E. Marketplace substitution of Atlantic salmon for Pacific salmon in Washington State detected by DNA barcoding[J]. Food Research International, 2012, 45(1): 388-393.
DOI URL |
| [27] |
Clark L F. The current status of DNA barcoding technology for species identification in fish value chains[J]. Food Policy, 2015, 54: 85-94.
DOI URL |
| [28] |
Chang C, Lin H, Ren Q, et al. DNA barcode identification of fish products in Taiwan: government-commissioned authentication cases[J]. Food Control, 2016, 66: 38-43.
DOI URL |
| [29] |
Zahn R J, Silva A J, Hellberg R S. Development of a DNA mini-barcoding protocol targeting COⅠfor the identification of elasmobranch species in shark cartilage pills[J]. Food Control, 2020, 109: 106918.
DOI URL |
| [30] |
Sultana S, Ali M E, Hossain M A M, et al. Universal mini COⅠbarcode for the identification of fish species in processed products[J]. Food Research International, 2018, 105: 19-28.
DOI URL |
| [31] |
Xiong X, Yuan F, Huang M, et al. DNA barcoding revealed mislabeling and potential health concerns with roasted fish products sold across China[J]. Journal of Food Protection, 2019, 82(7): 1200-1209.
DOI PMID |
| [32] |
Shedko S V, Miroshnichenko I L, Nemkova G A. Phylogeny of salmonids (Salmoniformes: Salmonidae) and its molecular dating: analysis of nuclear RAG1 gene[J]. Russian Journal of Genetics, 2012, 48: 575-579.
DOI URL |
| [33] |
Esin E V, Markevich G N. Evolution of the charrs, genus Salvelinus (Salmonidae). 1. origins and expansion of the species[J]. Journal of Ichthyology, 2018, 58(2): 187-203.
DOI |
| [34] | 李瑶瑶, 刘云国, 刘凌霄, 等. 鲑科鱼类线粒体全基因组序列结构特征及其系统发育信息分析[J]. 烟台大学学报(自然科学与工程版), 2016, 29(4):271-279. |
| [35] | 孙毅. 基于线粒体基因COX1、Cyt b和ND4的鲑科鱼类的系统发育[J]. 畜牧与饲料科学, 2015, 36(9):9-17. |
| [36] |
Mariani S, Griffiths A M, Velasco A, et al. Low mislabeling rates indicate marked improvements in Eusropean seafood market operations[J]. Frontiers in Ecology and the Environment, 2015, 13(10):536-540.
DOI URL |
| [37] |
Tang Q, Luo Q I, Duan Q, et al. DNA barcode identification of fish products from Guiyang markets in southwestern People’s Republic of China[J]. Journal of Food Protection, 2022, 85(4): 583-590.
DOI URL |
| [38] |
Cordes J F, Stephens M R, Blumberg M A, et al. Identifying introgressive hybridization in native populations of California golden trout based on molecular markers[J]. Transactions of the American Fisheries Society, 2006, 135(1): 110-128.
DOI URL |
| [39] |
Stephens M R, Clipperton N W, May B. Subspecies-informative SNP assays for evaluating introgression between native golden trout and introduced rainbow trout[J]. Molecular Ecology Resources, 2009, 9(1): 339-343.
DOI PMID |
| [40] |
Maxime V. The physiology of triploid fish: current knowledge and comparisons with diploid fish[J]. Fish and Fisheries, 2008, 9(1): 67-78.
DOI URL |
| [41] |
Manor M L, Weber G M, Cleveland B M, et al. Expression of genes associated with fatty acid metabolism during maturation in diploid and triploid female rainbow trout[J]. Aquaculture, 2015, 435: 178-186.
DOI URL |
| [42] | 王庆龙. 金鳟和虹鳟繁殖与育种关键技术研究[D]. 青岛: 中国海洋大学, 2013. |
| [1] | WANG Xiaoliang, CAO Huan, WANG Shu, LÜ Xiaonan, WANG Jingbo, ZHANG Wen, WANG Peng, XU Lipu. Morphological characteristic and molecular identification of Myxobolus wulii found in dorsolateral muscle and hepatopancreas of diseased goldfish(Carassius auratus) [J]. Journal of Fisheries Research, 2024, 46(1): 85-91. |
| [2] | . Analysis of the genetic diversity and sequence variation of mitochondrial COⅠ gene from three populations of Macrobrachium rosenbergii [J]. Journal of Fisheries Research, 2023, 45(1): 8-13. |
| [3] | Peng-yun WANG. Morphological characteristics and molecular phylogenetic analysis of the cultured oyster from the Houhai bay in Putian [J]. JOURNAL OF FUJIAN FISHERIES, 2013, 35(2): 100-105. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||