Journal of Fisheries Research ›› 2023, Vol. 45 ›› Issue (5): 415-426.DOI: 10.14012/j.cnki.fjsc.2023.05.001
• Research Paper • Next Articles
LI Chenrui( ), XU Rongliang, ZHANG Xiaoxu, CHEN Dongping, QIU Yuyang, CHEN Weiqin, QIN Yingxue*(
), XU Rongliang, ZHANG Xiaoxu, CHEN Dongping, QIU Yuyang, CHEN Weiqin, QIN Yingxue*( )
)
Received:2023-03-08
Online:2023-10-25
Published:2023-10-07
李辰睿( ), 徐荣亮, 张小旭, 陈东萍, 邱于洋, 陈伟琴, 覃映雪*(
), 徐荣亮, 张小旭, 陈东萍, 邱于洋, 陈伟琴, 覃映雪*( )
)
通讯作者:
覃映雪(1976—),女,教授,硕士生导师,研究方向为水生动物病原微生物。E-mail:作者简介:李辰睿(2001—),女,本科,研究方向为水生动物医学。E-mail:1091222815@qq.com
基金资助:CLC Number:
LI Chenrui, XU Rongliang, ZHANG Xiaoxu, CHEN Dongping, QIU Yuyang, CHEN Weiqin, QIN Yingxue. Genome analysis of pathogenic Vibrio harveyi strain TS-628[J]. Journal of Fisheries Research, 2023, 45(5): 415-426.
李辰睿, 徐荣亮, 张小旭, 陈东萍, 邱于洋, 陈伟琴, 覃映雪. 病原哈维氏弧菌(Vibrio harveyi)TS-628菌株基因组分析[J]. 渔业研究, 2023, 45(5): 415-426.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.hyyysci.com/EN/10.14012/j.cnki.fjsc.2023.05.001

Fig.1 GC% and sequencing depth profiles of strains TS-628 Notes:The abscissa of the scatter chart is the GC content,and the ordinate is the read coverage depth;the histogram at right and up sides shows the frequency distribution of sliding window of GC content and sequencing depth,respectively.
| 样品名称 Sample name | 染色体数量 Chromosome no. | 质粒数量 Plasmid no. | 基因长度/bp Genome size | GC含量/% GC content |
|---|---|---|---|---|
| 哈维氏弧菌 V.harveyi | 2 | 2 | 6 151 198 | 44.68 |
Tab.1 The genome assembly results of strains TS-628
| 样品名称 Sample name | 染色体数量 Chromosome no. | 质粒数量 Plasmid no. | 基因长度/bp Genome size | GC含量/% GC content |
|---|---|---|---|---|
| 哈维氏弧菌 V.harveyi | 2 | 2 | 6 151 198 | 44.68 |
| 程序集 Assembly ID | 序列号 Sequence ID | 样品名称 Sample name | 长度/bp Length | GC含量/% GC content | 组成信息 Bases statistics | 序列类型 Sequence type |
|---|---|---|---|---|---|---|
| 染色体1 Chromosome 1 | Scaffold 1 | V.harveyi | 3 574 667 | 44.95 | A:980761;T:986964; G:80357;C:803371 | Chromosome |
| 染色体2 Chromosome 2 | Scaffold 2 | V.harveyi | 2 292 642 | 44.76 | A:633107;T:633402; G:514578;C:511555 | Chromosome |
| 质粒A Plasmid A | Scaffold 3 | V.harveyi | 217 554 | 39.41 | A:61456;T:70356; G:43418;C:42324 | Known-Plasmid |
| 质粒B Plasmid B | Scaffold 4 | V.harveyi | 66 335 | 44.56 | A:17821;T:18954; G:15310;C:14250 | New-Plasmid |
Tab.2 The genome assembly details of strains TS-628
| 程序集 Assembly ID | 序列号 Sequence ID | 样品名称 Sample name | 长度/bp Length | GC含量/% GC content | 组成信息 Bases statistics | 序列类型 Sequence type |
|---|---|---|---|---|---|---|
| 染色体1 Chromosome 1 | Scaffold 1 | V.harveyi | 3 574 667 | 44.95 | A:980761;T:986964; G:80357;C:803371 | Chromosome |
| 染色体2 Chromosome 2 | Scaffold 2 | V.harveyi | 2 292 642 | 44.76 | A:633107;T:633402; G:514578;C:511555 | Chromosome |
| 质粒A Plasmid A | Scaffold 3 | V.harveyi | 217 554 | 39.41 | A:61456;T:70356; G:43418;C:42324 | Known-Plasmid |
| 质粒B Plasmid B | Scaffold 4 | V.harveyi | 66 335 | 44.56 | A:17821;T:18954; G:15310;C:14250 | New-Plasmid |
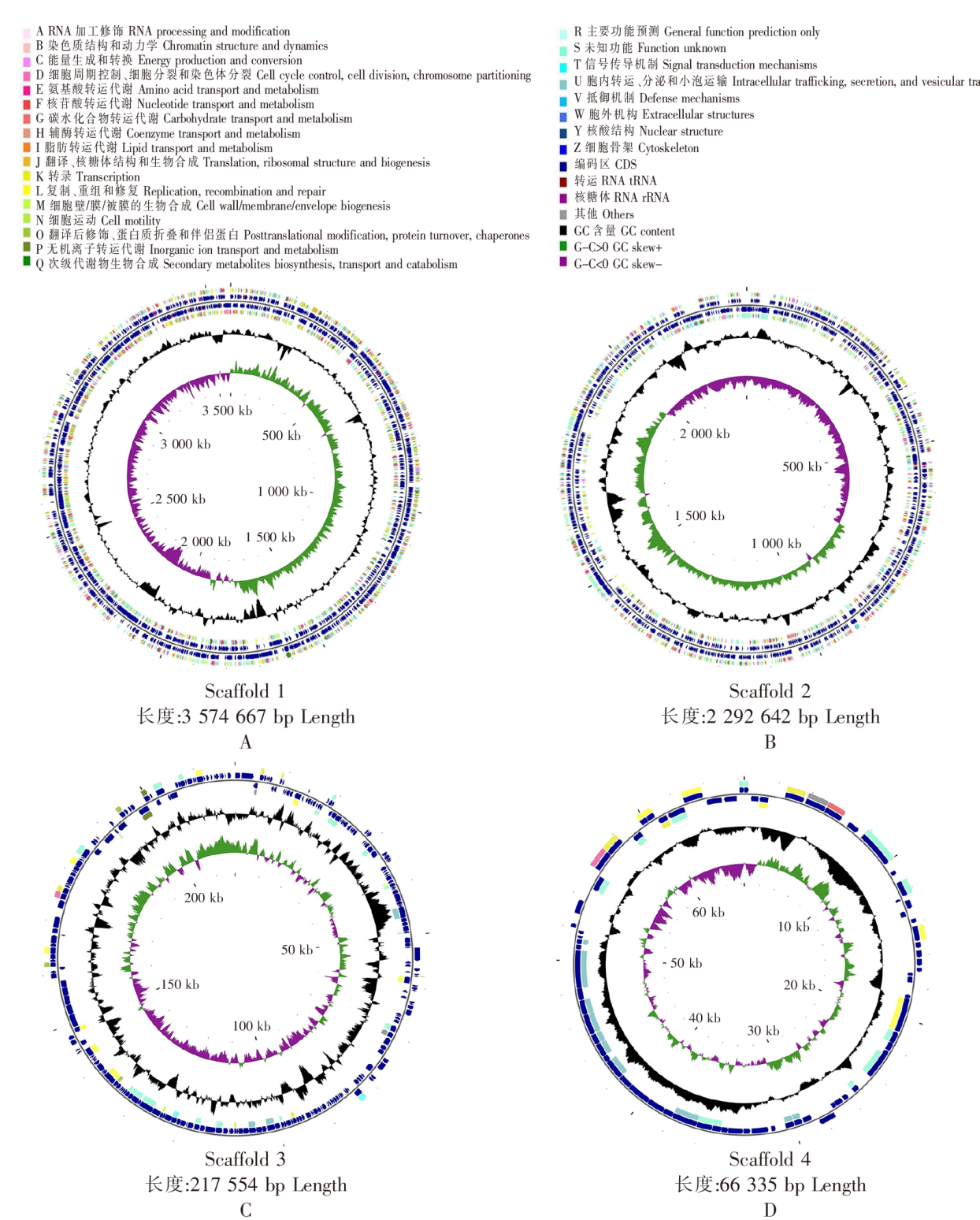
Fig.2 The CGView map of genome Notes:Figture 2A is chromosome 1;Figture 2B is chromosome 2;Figture 2C is plasmid A;Figture 2D is plasmid B.From the inner circle to the outer circle,the first circle represents the total length of the genome;the second circle represents the GC skeleton,where green represents G-C>0,while purple represents the opposite,the intersection of green and purple represents the start point and end point of replication,respectively;the black in the third circle indicates the GC content,and the protruding part of the peak indicates that the GC content in this region is higher than the average GC content of the whole genome,while the opposite is true inward;the fourth and fifth circles respectively represent genes on the positive and negative chains,including tRNA,rRNA,CDS and other genes.
| 样品名称 Sample name | 类别1 Level 1 | 类别2 Level 2 | 基因数量 Gene no. | 类别1 Level 1 | 类别2 Level 2 | 基因数量 Gene no. | |
|---|---|---|---|---|---|---|---|
| 防御性毒力因子 | 应激蛋白 | 13 | 攻击性毒力因子 | 入侵类 | 13 | ||
| 哈维氏弧菌 V.harveyi | 非特异性毒力因子 | 铁吸收系统 | 130 | 防御性毒力因子 | 相位变异 | 9 | |
| 防御性毒力因子 | 抗吞噬类 | 36 | 防御性毒力因子 | 抗血清 | 11 | ||
| 毒力调控相关基因 | 调控相关 | 35 | 非特异性毒力因子 | 镁吸收系统 | 4 | ||
| 攻击性毒力因子 | 分泌系统 | 107 | 防御性毒力因子 | 补体蛋白酶 | 2 | ||
| 攻击性毒力因子 | 附着类 | 132 | 攻击性毒力因子 | 毒素 | 30 |
Tab.3 The prediction and classification of virulence genes
| 样品名称 Sample name | 类别1 Level 1 | 类别2 Level 2 | 基因数量 Gene no. | 类别1 Level 1 | 类别2 Level 2 | 基因数量 Gene no. | |
|---|---|---|---|---|---|---|---|
| 防御性毒力因子 | 应激蛋白 | 13 | 攻击性毒力因子 | 入侵类 | 13 | ||
| 哈维氏弧菌 V.harveyi | 非特异性毒力因子 | 铁吸收系统 | 130 | 防御性毒力因子 | 相位变异 | 9 | |
| 防御性毒力因子 | 抗吞噬类 | 36 | 防御性毒力因子 | 抗血清 | 11 | ||
| 毒力调控相关基因 | 调控相关 | 35 | 非特异性毒力因子 | 镁吸收系统 | 4 | ||
| 攻击性毒力因子 | 分泌系统 | 107 | 防御性毒力因子 | 补体蛋白酶 | 2 | ||
| 攻击性毒力因子 | 附着类 | 132 | 攻击性毒力因子 | 毒素 | 30 |
| 样品名称 Sample name | 类型1 Type Ⅰ | 类型2 Type Ⅱ | 类型3 Type Ⅲ | 类型4 Type Ⅳ | 类型5 Type Ⅴ | 类型6 Type Ⅵ | Sec-SRP型 Sec-SRP | Tat型 Tat | 基因总数 Total gene no. |
|---|---|---|---|---|---|---|---|---|---|
| V.harveyi | 1 | 12 | 17 | 1 | 0 | 7 | 13 | 3 | 54 |
Tab.4 The classification and statistics of secretory system
| 样品名称 Sample name | 类型1 Type Ⅰ | 类型2 Type Ⅱ | 类型3 Type Ⅲ | 类型4 Type Ⅳ | 类型5 Type Ⅴ | 类型6 Type Ⅵ | Sec-SRP型 Sec-SRP | Tat型 Tat | 基因总数 Total gene no. |
|---|---|---|---|---|---|---|---|---|---|
| V.harveyi | 1 | 12 | 17 | 1 | 0 | 7 | 13 | 3 | 54 |

Fig.5 The secretory pathway Notes:The secretory system pathway diagram is a graphical representation of all secretory systems in each sample.The red box marks the genes related to the secretion system predicted in the sample.
| 样品名称 Sample name | 调控基因数量 Gene no.of regulator | 传感基因数量 Gene no.of sensor | 混合基因数量 Gene no.of hybrid |
|---|---|---|---|
| V.harveyi | 149 | 63 | 30 |
Tab.5 The statistics of two-component regulation classification
| 样品名称 Sample name | 调控基因数量 Gene no.of regulator | 传感基因数量 Gene no.of sensor | 混合基因数量 Gene no.of hybrid |
|---|---|---|---|
| V.harveyi | 149 | 63 | 30 |
| 菌株 Strains | 菌株来源 Source | 基因 长度/bp Genome length | GC 含量/% GC content | 染色体 数量 Chromosome no. | 质粒 数量 Plasmid no. | 主要毒力基因种类 Major virulence gene species |
|---|---|---|---|---|---|---|
| TS-628(PRJNA874896) | 青石斑鱼 | 6 151 198 | 44.68 | 2 | 2 | 附着类、铁摄取、分泌类 |
| QT-520(PRJNA340970) | 卵形鲳鲹 | 6 070 846 | 45.05 | 2 | 3 | 附着类、抗吞噬类、毒素 |
| 345(PRJNA418027) | 龙胆石斑 | 6 185 822 | 44.76 | 2 | 2 | 附着类、铁摄取、运动性、分泌类 |
Tab.6 Comparative analysis table of the main virulence factors of 3 strains of V.harveyi
| 菌株 Strains | 菌株来源 Source | 基因 长度/bp Genome length | GC 含量/% GC content | 染色体 数量 Chromosome no. | 质粒 数量 Plasmid no. | 主要毒力基因种类 Major virulence gene species |
|---|---|---|---|---|---|---|
| TS-628(PRJNA874896) | 青石斑鱼 | 6 151 198 | 44.68 | 2 | 2 | 附着类、铁摄取、分泌类 |
| QT-520(PRJNA340970) | 卵形鲳鲹 | 6 070 846 | 45.05 | 2 | 3 | 附着类、抗吞噬类、毒素 |
| 345(PRJNA418027) | 龙胆石斑 | 6 185 822 | 44.76 | 2 | 2 | 附着类、铁摄取、运动性、分泌类 |
| [1] |
Zorrilla I, Arijo S, Chabrillon M, et al. Vibrio species isolated from diseased farmed sole,Solea senegalensis(Kaup),and evaluation of the potential virulence role of their extracellular products[J]. Journal of Fish Diseases, 2003, 26(2):103-108.
PMID |
| [2] |
Karunasagar I, Pai R, Malathi G R, et al. Mass mortality of Penaeus monodon larvae due to antibiotic-resistant Vibrio harveyi infection[J]. Aquaculture, 1994, 128(3-4):203-209.
DOI URL |
| [3] |
Nishimori E, Hasegawa O, Numata T, et al. Vibrio carchariae causes mass mortalities in Japanese abalone,Sulculus diversicolor supratexta[J]. Fish Pathology, 1998, 33(5):495-502.
DOI URL |
| [4] |
Luna G M, Bongiorni L, Gili C, et al. Vibrio harveyi as a causative agent of the white syndrome in tropical stony corals[J]. Environmental Microbiology Reports, 2010, 2(1):120-127.
DOI URL |
| [5] |
Robertson P A W, Xu H S, Austin B. An enzyme-linked immunosorbent assay (ELISA) for the detection of Vibrio harveyi in penaeid shrimp and water[J]. Journal of Microbiological Methods, 1998, 34(1):31-39.
DOI URL |
| [6] |
Liu P C, Lee K K, Chen S N. Pathogenicity of different isolates of Vibrio harveyi in tiger prawn,Penaeus monodon[J]. Letters in Applied Microbiology, 1996, 22(6):413-416.
DOI URL |
| [7] | 黄辉. 哈维弧菌表面抗原的克隆和真核表达[D]. 武汉: 华中农业大学, 2010. |
| [8] | 金珊, 王国良, 赵青松, 等. 海水网箱养殖大黄鱼弧菌病的流行病学研究[J]. 水产科学, 2005, 24(1):17-19. |
| [9] |
Kraxberger-Beatty T, McGarey D J, Grier H J, et al. Vibrio harveyi,an opportunistic pathogen of common snook,Centropomus undecimalis(Bloch),held in captivity[J]. Journal of Fish Diseases, 1990, 13(6):557-560.
DOI URL |
| [10] | 陈献稿, 吴淑勤, 石存斌, 等. 斜带石斑鱼病原菌(哈维氏弧菌)的分离与鉴定[J]. 中国水产科学, 2004, 11(4):313-317. |
| [11] |
Soffientino B, Gwaltney T, Nelson D R, et al. Infectious necrotizing enteritis and mortality caused by Vibrio carchariae in summer flounder Paralichthys dentatus during intensive culture[J]. Diseases of Aquatic Organisms, 1999, 38(3):201-210.
PMID |
| [12] | 徐晓丽, 邵蓬, 李贺密, 等. 半滑舌鳎体表溃疡病病原的分离鉴定[J]. 中国海洋大学学报(自然科学版), 2015, 45(11):29-35. |
| [13] | 张宏伟, 付建荣, 苏东, 等. 弧菌科细菌致急性腹泻的流行病学调查[J]. 中国医学检验杂志, 1996, 19(1):45. |
| [14] |
Austin B, Zhang X H. Vibrio harveyi:a significant pathogen of marine vertebrates and invertebrates[J]. Letters in Applied Microbiology, 2006, 43(2):119-124.
PMID |
| [15] |
Natrah F M, Ruwandeepika H A, Pawar S, et al. Regulation of virulence factors by quorum sensing in Vibrio harveyi[J]. Veterinary Microbiology, 2011, 154(1-2):124-129.
DOI PMID |
| [16] | Zhang X H, He X X, Austin B. Vibrio harveyi:a serious pathogen of fish and invertebrates in mariculture[J]. Marine Life Science & Technology, 2020, 2(3):231-245. |
| [17] |
Strauss E J, Falkow S. Microbial pathogenesis:genomics and beyond[J]. Science, 1997, 276(5313):707-712.
PMID |
| [18] | Qin Y X, Wang J, Su Y Q, et al. Studies on the pathogenic bacterium of ulcer disease in Epinephelus awoara[J]. Acta Oceanologica Sinica, 2006, 25(1):154-159. |
| [19] |
Besemer J, Lomsadze A, Borodovsky M. GeneMarkS:a self-training method for prediction of gene starts in microbial genomes.Implications for finding sequence motifs in regulatory regions[J]. Nucleic Acids Research, 2001, 29(12):2607-2618.
DOI PMID |
| [20] |
Grant R J, Stothard P. The CGView Server:a comparative genomics tool for circular genomes[J]. Nucleic Acids Research, 2008, 36(Suppl_2):W181-W184.
DOI URL |
| [21] |
Jehl M A, Arnold R, Rattei T. Effective-a database of predicted secreted bacterial proteins[J]. Nucleic Acids Research, 2011, 39(Suppl-1):D591-D595.
DOI URL |
| [22] | 徐先栋. 华南海水养殖地区哈氏弧菌分子流行病学特征及其毒力菌株基因组分析[D]. 海口: 海南大学, 2017. |
| [23] | Wang Z, Hervey W J Ⅳ, Kim S, et al. Complete genome sequence of the bioluminescent marine bacterium Vibrio harveyi ATCC 33843 (392 [MAV])[J]. Genome Announcements, 2015, 3(1):e01493-e01414. |
| [24] |
Moriyama E N, Powell J R. Gene length and codon usage bias in Drosophila melanogaster,Saccharomyces cerevisiae and Escherichia coli[J]. Nucleic Acids Research, 1998, 26(13):3188-3193.
PMID |
| [25] |
Tao Y, Zhou K X, Xie L Y, et al. Emerging coexistence of three PMQR genes on a multiple resistance plasmid with a new surrounding genetic structure of qnrS2 in E.coli in China[J]. Antimicrobial Resistance and Infection Control, 2020, 9(1):52-59.
DOI |
| [26] |
Levinson B L, Kasyan K J, Chiu S S, et al. Identifieation of β-exotoxin production,plasmids eneoding β-exotoxin and a new exotoxin in Bacillus thuringiensis by using high-performance liquid chromatography[J]. Journal of Bacteriology, 1990, 172(6):3172-3179.
PMID |
| [27] |
Singer J T, Earley S. Identification of polypeptides encoded by cloned pJM1 iron uptake DNA isolated from Vibrio anguillarum 775[J]. Journal of Bacteriology, 1989, 171 (5):2293-2302.
PMID |
| [28] |
Virdle C, Goldlust K, Djermoun S, et al. Plasmid transfer by conjugation in gram-negative bacteria:from the cellular to the community level[J]. Genes, 2020, 11(11):1239-1271.
DOI URL |
| [29] |
Munro J, Oakey J, Bromage E, et al. Experimental bacteriophage-mediated virulence in strains of Vibrio harveyi[J]. Diseases of Aquatic Organisms, 2003, 54(3):187-194.
DOI URL |
| [30] |
Prasad S, Mingrino R, Kaukinen K, et al. Inflammatory processes have differential effects on claudins 2,3 and 4 in colonic epithelial cells[J]. Laboratory Investigation, 2005, 85(9):1139-1162.
DOI PMID |
| [31] |
Bai F, Pang L, Qi Z, et al. Distribution of five Vibrio virulence-related genes among Vibrio harveyi isolates[J]. The Journal of General and Applied Microbiology, 2008, 54(1):71-78.
DOI URL |
| [32] |
Tu Z, Li H, Zhang X, et al. Complete genome sequence and comparative genomics of the golden pompano (Trachinotus ovatus) pathogen,Vibrio harveyi strain QT520[J]. PeerJ, 2017, 5:e4127.
DOI URL |
| [33] |
Deng Y Q, Xu H D, Su Y L, et al. Horizontal gene transfer contributes to virulence and antibiotic resistance of Vibrio harveyi 345 based on complete genome sequence analysis[J]. BMC Genomics, 2019, 20(1):1-19.
DOI |
| [34] |
Nakasone N, Iwanaga M. Pili of a Vibrio parahaemolyticus strain as a possible colonization factor[J]. Infection and Immunity, 1990, 58 (1):61-69.
DOI PMID |
| [35] | Sperandio V, Girón J A, Silveira W D, et al. The OmpU outer membrane protein,a potential adherence factor of Vibrio cholerae[J]. Infection & Immunity, 1995, 63(11):4433-4438. |
| [36] |
Nagayama K, Oguchi T, Arita M, et al. Purification and characterization of a cell-associated hemagglutinin of Vibrio parahaemolyticus[J]. Infection and Immunity, 1995, 63 (5):1987-1992.
DOI PMID |
| [37] | 张莹, 张文莉, 陈小贝, 等. 细菌产铁载体的结构、功能及其研究进展[J]. 中国卫生检验杂志, 2012, 22(9):2249-2251. |
| [38] | 蔡明辉, 吕春梅, 张晓兰, 等. 革兰氏阳性细菌铁摄取机制的研究进展[J]. 国际免疫学杂志, 2016, 39(3):245-249. |
| [39] |
Litwin C M, Calderwood S B. Role of iron in regulation of virulence genes[J]. Clinical Microbiology Reviews, 1993, 6(2):137-149.
DOI PMID |
| [40] |
Naka H, Actis L A, Crosa J H. The anguibactin biosynthesis and transport genes are encoded in the chromosome of Vibrio harveyi:a possible evolutionary origin for the pJM1 plasmid-encoded system of Vibrio anguillarum?[J]. MicrobiologyOpen, 2013, 2(1):182-194.
DOI URL |
| [41] |
Desvaux M, Hébraud M, Talon R, et al. Secretion and subcellular localizations of bacterial proteins:a semantic awareness issue[J]. Trends in Microbiology, 2009, 17(4):139-145.
DOI URL |
| [42] | 白镜塬. 哈维氏弧菌GDH11385的紫外诱变及其诱变菌株的比较基因组学分析[D]. 海口: 海南大学, 2018. |
| [43] |
Huang W, Li L, Myers J R, et al. ART:a next-generation sequencing read simulator[J]. Bioinformatics, 2012, 28(4):593-594.
DOI URL |
| [44] |
Espinoza-Valles I, Soto-Rodriguez S, Edwards R A, et al. Draft genome sequence of the shrimp pathogen Vibrio harveyi CAIM 1792[J]. Journal of Bacteriology, 2012, 194(8):2104-2104.
DOI PMID |
| [45] |
Dalbey R E, Kuhn A. Protein traffic in Gram-negative bacteria——how exported and secreted proteins find their way[J]. FEMS Microbiology Reviews, 2012, 36(6):1023-1045.
DOI PMID |
| [46] |
Korotkov K V, Gray M D, Kreger A, et al. Calcium is essential for the major pseudopilin in the type 2 secretion system[J]. Journal of Biological Chemistry, 2009, 284(38):25466-25470.
DOI PMID |
| [47] |
Sandkvist M, Michel L O, Hough L P, et al. General secretion pathway (eps) genes required for toxin secretion and outer membrane biogenesis in Vibrio cholerae[J]. Journal of Bacteriology, 1997, 179(22):6994-7003.
PMID |
| [48] |
Tauschek M, Gorrell R J, Strugnell R A, et al. Identification of a protein secretory pathway for the secretion of heat-labile enterotoxin by an enterotoxigenic strain of Escherichia coli[J]. Proceedings of the National Academy of Sciences of the United States of America, 2002, 99(10):7066-7071.
DOI PMID |
| [49] |
Hirst T R, Holmgren J. Conformation of protein secreted across bacterial outer membranes:a study of enterotoxin translocation from Vibrio cholerae[J]. Proceedings of the National Academy of Sciences of the United States of America, 1987, 84(21):7418-7422.
PMID |
| [50] | Sandkvist M. Type Ⅱ secretion and pathogenesis[J]. Infection & Immunity, 2001, 69(6):3523-3535. |
| [51] | McCallum M, Burrows L L, Howell P L. The dynamic structures of the type Ⅳ Pilus[J]. Microbiology Spectrum, 2019, 7(2):113-128. |
| [52] | Darshanee Ruwandeepika H A, Karunasagar I, Bossier P, et al. Expression and quorum sensing regulation of type Ⅲ secretion system genes of Vibrio harveyi during infection of gnotobiotic brine shrimp[J]. PLoS One, 2015, 10(12):e0143935. |
| [53] |
Okada N, Matsuda S, Matsuyama J, et al. Presence of genes for type Ⅲ secretion system 2 in Vibrio mimicus strains[J]. BMC Microbiology, 2010, 10:302.
DOI |
| [54] |
Park K S, Ono T, Rokuda M, et al. Functional characterization of two type Ⅲ secretion systems of Vibrio parahaemolyticus[J]. Infection and Immunity, 2004, 72(11):6659-6665.
DOI URL |
| [55] |
Portaliou A G, Tsolis K C, Loos M S, et al. Type Ⅲ secretion:building and operating a remarkable nanomachine[J]. Trends in Biochemical Sciences, 2016, 41(2):175-189.
DOI PMID |
| [56] |
Morgado S, Vicente A C. Diversity and distribution of type Ⅵ secretion system gene clusters in bacterial plasmids[J]. Scientific Reports, 2022, 12(1):8249.
DOI PMID |
| [57] |
Cassan F D, Coniglio A, Amavizca E, et al. The Azospirillum brasilense type Ⅵ secretion system promotes cell aggregation,biocontrol protection against phytopathogens and attachment to the microalgae Chlorella sorokiniana[J]. Environmental Microbiology, 2021, 23(10):6257-6274.
DOI URL |
| [58] |
Li C F, Zhu L F, Wang D D, et al. T6SS secretes an LPS-binding effector to recruit OMVs for exploitative competition and horizontal gene transfer[J]. The ISME Journal, 2022, 16(2):500-510.
DOI URL |
| [1] | . Investigation of the bacterial diversity and analysis of environmental influencing factors in the Luoyuan Bay [J]. Journal of Fisheries Research, 2021, 43(4): 384-393. |
| [2] | . Research on the biofloc bacterial community structure during larval rearing of Litopenaeus vannamei using metagenome sequencing [J]. , 2015, 37(2): 91-97. |
| [3] | . Molecular characterization of six strains of Aeromonas hydrophila isolated from fish as well as detection and analysis of virulence genes [J]. , 2014, 36(2): 87-95. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||