弧菌属于细菌界(Bacteria)、变形菌门(Proteobacteria)、γ-变形菌纲 (Gammaproteobacteria)、弧菌目(Vibrionales)、弧菌科(Vibrionaceae)、弧菌属(Vibrio)[1-2]。弧菌科是弧菌目中唯一的科,包括以弧菌属为代表的8个属,共计159个种,主要分布在海洋与河口环境中,在淡水中偶尔也有分布,某些种类是水产动物与人类重要的病原菌,具有潜在的环境效应[3]。致病性弧菌如哈维氏弧菌、溶藻弧菌和副溶血弧菌等,是导致水产动物病害暴发的重要原因[4⇓-6];人们食用了遭弧菌污染的水产动物,会引发腹泻、腹痛、呕吐等食物中毒症状[7];伤口感染了创伤弧菌,会引发组织溃烂,导致组织坏死,甚至危及生命安全[8]。在一次我国13个地区2 980份生食动物性水产品样本调查中,副溶血性弧菌检出率达到14.7%,创伤弧菌检出率为3.5%[9],可见弧菌对人类的危害性不容忽视。
16S rDNA基因序列长约1.5 kb,具有高度的保守性和特异性,常被用于原核微生物系统发育学方面的研究 [10]。16S rDNA作为原核生物分类的重要分子标记,在细菌鉴定相关工作中起到重要作用[11⇓-13]。热休克蛋白60(HSP60)基因作为HSPs家族中最为保守的亚类,广泛存在于真核生物及原核生物中,发挥重要的调控作用[14]。促旋酶(Gyrase)的B亚单位(gyrB)普遍存在于细菌核酸中,该基因呈单拷贝,比16S rDNA的进化速度快,而且包含可变区和保守区,符合用于细菌系统发育靶基因的要求,有利于细菌鉴定及快速检测[15-16]。本研究从西沙永乐群岛海域的海水中分离弧菌菌株,并用多个基因引物进行序列扩增与比对分析,确定一种可靠的海洋弧菌分子鉴定方法,同时对西沙永乐群岛海域海洋弧菌的多样性进行初步调查与评价,旨在为相关方面的研究提供参考。
1 材料与方法
1.1 试验材料
海水样品取自中国南海西沙永乐群岛7个岛礁(晋卿岛、石屿、银屿、全富岛、鸭公岛、甘泉岛、羚羊礁筐仔沙洲)近岸水域,每个岛礁设两个位点,每个取样位点的海水样品取15 L,用0.22 μL微孔滤膜进行抽滤,滤膜放于离心管中,置于-20℃冰箱中保存并带回实验室。
1.2 试验方法
1.2.1 菌株分离与纯化
配制TCBS琼脂培养基,于121℃灭菌20 min,冷却至50℃后倒入一次性培养皿中,培养基厚度约6 mm。取出微孔滤膜,浸泡于15 mLPBS缓冲液中,进行超声波振荡10 min,取100 μL涂布于TCBS平板上,然后置于30℃生化培养箱中培养24 h;另取100 μL接种到已灭菌的TCBS液体培养基中,置于30℃生化培养箱中培养24 h后,取100 μL涂布于TCBS平板上,再置于30℃生化培养箱中培养24 h。挑取不同形态特征的单菌落至TCBS平板中,利用划线分离法进一步纯化,置于30℃生化培养箱中培养24 h,重复划线分离纯化2次,获得纯种菌株。
1.2.2 菌株的鉴定
1)形态学及显微镜观察
观察菌落状态,挑选纯种菌株进行革兰氏染色,在显微镜下观察形态特征。
2)16S rDNA基因扩增及检测
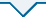
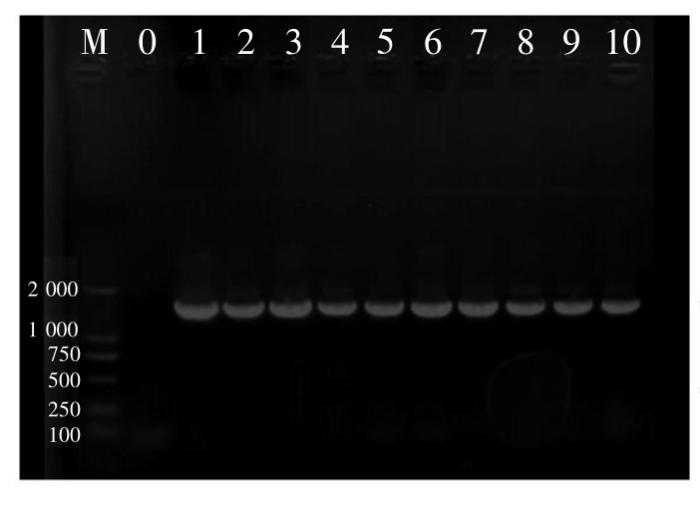
挑取纯化的单菌落至20 μL无菌水中作为扩增模板。PCR扩增反应体系(25 μL):2×F8 FastLong PCR MasterMIX 12.5 μL,引物16S-F和16S-R各1 μL,模板1 μL,无菌超纯水9.5 μL。反应条件:94℃预变性4 min,94℃变性10 s,55℃退火15 s;72℃延伸15 s,34个循环,72℃温育5 min。PCR产物用含Goldview的1%琼脂糖凝胶进行电泳检测,电压110 V,时间30 min。选取大小约1 500 bp的PCR产物送样至深圳华大基因股份有限公司进行测序,将测序得到的菌株16S rDNA序列在GenBank中进行BLAST比对,若有相似度高的序列信息则进行下载,并用MEGA 6.0软件构建系统发育树。
3)HSP60基因扩增及检测
挑取纯化的单菌落至20 μL无菌水中作为扩增模板。PCR反应体系(25 μL):2×F8 FastLong PCR MasterMIX 12.5 μL,引物HSP60-F和HSP60-R各1 μL,模板1 μL,无菌超纯水9.5 μL。反应条件:94℃预变性4 min,94℃变性10 s,50℃退火15 s,72℃延伸10 s,30个循环;72℃ 温育5 min。PCR产物用含Goldview的1%琼脂糖凝胶进行电泳检测,电压90 V,时间30 min。选取大小约600 bp的PCR产物送样至深圳华大基因股份有限公司进行测序,将测序得到的菌株HSP60序列在GenBank中进行BLAST比对,若有相似度高的序列信息则进行下载,并用MEGA 6.0软件构建系统发育树。
4)gyrB基因扩增及检测
设计3对gyrB引物,见表1。挑取纯化的单菌落至20 μL无菌水中作为扩增模板,PCR反应体系(25 μL):2×F8 FastLong PCR MasterMIX 12.5 μL,正向引物和反向引物各1 μL,模板1 μL,无菌超纯水9.5 μL。反应条件:94℃预变性3 min,94℃变性10 s,55℃退火15 s,72℃延伸10 s,34个循环;72℃温育5 min。PCR产物用含Goldview的1%琼脂糖凝胶进行电泳检测,电压110 V,时间30 min。选取大小约1 200 bp的PCR产物送样至深圳华大基因股份有限公司进行测序。将测序得到的菌株gyrB序列在GenBank中进行BLAST比对,若有相似度高的序列信息则进行下载,并用MEGA 6.0软件构建系统发育树。
表1 引物序列
Tab.1
| 目标基因 Target genes | 引物 Primers | 引物序列(5’-3’) Sequences(5’-3’) | 产物大小/bp Amplification size |
|---|---|---|---|
| 16S rDNA | 16S-F(通用引物) | AGAGTTTGATCCTGGCTCAG | 1 400~1 500 |
| 16S-R(通用引物) | GGTTACCTTGTTACGACTT | ||
| HSP60 | HSP60-F[4] | ACAACAGCAACGGTACTAGC | 500~550 |
| HSP60-R[4] | CAACTTTCACGATGCCAC | ||
| gyrB | gyrB1-F[17] | TCCGGCGGTCTGCACGGCGT | 1 200~1 300 |
| gyrB1-R[17] | TTGTCCGGGTTGTACTCGTC | ||
| gyrB2-F[5] | AAGTCATCATGACCGTTCTGCAYGCNGGNGGNAARTTYGA | ||
| gyrB2-R[5] | AGCAGGGTACGGATGTGGCGAGCCRTCNACRTCNGCRTCNGTCAT | ||
| gyrB3-F[5] | GAAGTTATCATGACGGTACTTC | ||
| gyrB3-R[5] | CCTTTACGACGAGTCATTTC |
注:序列中“R”“N”为简并碱基,“R”对应的正常碱基为A/G,“N”对应的正常碱基为A/T/C/G。
Notes:In the sequence,“R” and “N” were degenerate bases,“R” corresponded to A/G,and “N” corresponded to A/T/C/G.
2 结果
2.1 菌株的形态学分析
根据不同的菌落形态对菌株进行挑选,共挑选出菌株47株,其中晋卿岛10株、石屿5株、银屿5株、全富岛6株、鸭公岛9株、甘泉岛7株、筐仔沙洲5株。47株菌株全为革兰氏阴性菌,在显微镜下形态为短小的杆状、弯曲的弧状或者S形(表2)。
表2 菌株的菌落形态及颜色
Tab.2
| 菌落形态 Colony morphology | 图示 Figure | 代表菌株 Representative strains |
|---|---|---|
| 扁平圆形,菌落干燥,表面光滑,边缘平整 The colonnies were flat,round and dry,and had smooth surface and flat edge |  | YG1-4(Y)、YG1-5(Y)、GQ2-2(Y)、 GQ2-3(Y) |
| 扁平圆形,菌落湿润,表面光滑,边缘平整 The colonnies were flat,round and moist,and had smooth surface and flat edge |  | LY2-4(Y)、QF2-1(Y)、QF2-2(Y)、 JQ2-1(G)、GQ1-2(Y) |
| 不规则圆形,菌落湿润,表面光滑,边缘平整,菌落中间有数个小突起 The colonnies were irregular circle and moist,and had smooth surface and flat edge.There were several small protuberances in the middle of the colonnies | 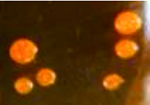 | YY1-7(Y)、JQ2-1(G)、QF2-4(Y) |
| 形状不规则,表面突起,菌落湿润,扩散状,边缘皱褶 The colonnies were irregular shape,moist and diffused,and had protuberance surface and wrinkle edge |  | GQ1-1(G)、GQ2-4(G)、SY2-2(G)、 SY2-6(G)、QF1-1(G)、QF1-2(G)、 QF2-1(G) |
| 形状不规则,菌落扁平,稍微扩散,湿润 The colonnies were irregular shape,moist and slightly diffused,and had smooth surface | 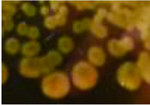 | LY2-3(G)、YG2-4(G)、YG2-5(G)、 QF2-5(G)、GQ1-5(G) |
| 圆形,菌落湿润,中间凸起,凸起为墨绿色,边缘为青白色 The colonnies were round and moist.The colonies bulge in the middle.Middle of colonies were dark green while edge of colonies were light green |  | LY2-1(G)、LY2-2(G) |
注:括号内字母表示菌株的菌落颜色,Y表示黄色,G表示绿色。
Notes:The letters in parentheses indicated the colony color of the strain.Y indicated yellow,G indicated green.
2.2 菌株的分子鉴定
2.2.1 16S rDNA鉴定
图1
图2
2.2.2 HSP60鉴定
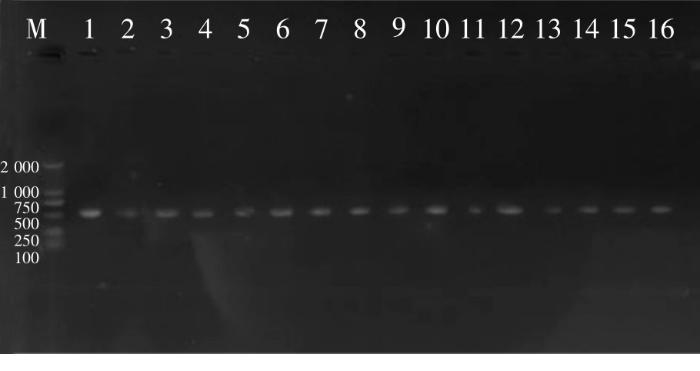
利用HSP60引物,47株菌株均能扩增出目标条带,部分菌株扩增条带见图3。47株菌株的比对结果均显示,与实验菌株相似度高于90%的菌株只有Vibrio neocaledonicus和Vibrio alginolyticus,无法构建系统发育树,因此利用HSP60基因不能成功鉴定弧菌到种。
图3
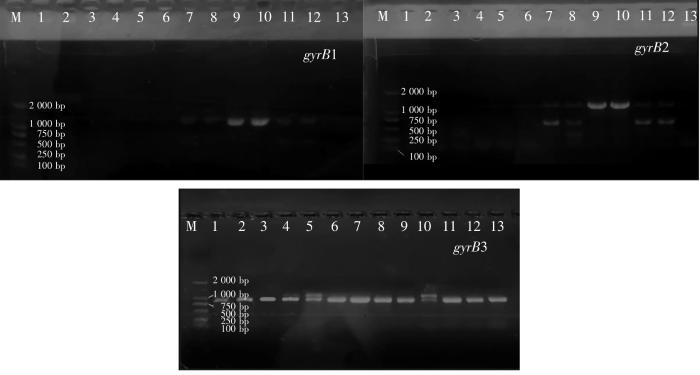
2.2.3 gyrB鉴定
图4
图5
通过gyrB3序列的系统发育分析,47株菌株分属于7个种:Vibrio alginolyticus、Vibrio harveyi、Vibrio mediterranei、Vibrio owensii、Vibrio coralliilyticus、Vibrio fortis、Vibrio variabilis。
2.3 不同岛礁海域的弧菌多样性分析
经gyrB3序列的分子鉴定,鸭公岛海域分离鉴定到5种弧菌,弧菌多样性最高;甘泉岛海域次之,分离鉴定到4种弧菌;石屿和羚羊礁筐仔沙洲的弧菌多样性最低,均只分离鉴定到2种弧菌。溶藻弧菌在7个岛礁海域中均被分离到,强壮弧菌只在银屿海域中被分离到(表3)。
表3 不同岛礁海域的弧菌多样性
Tab.3
| 地点 Locale | 菌株数量 Number of strains | 种 Species | 种的数量 Number of species |
|---|---|---|---|
| 晋卿岛 Jinqing Island | 10 | Vibrio alginolyticus、Vibrio harveyi、Vibrio mediterranei | 3 |
| 石屿 Shiyu Island | 5 | Vibrio alginolyticus、Vibrio coralliilyticus | 2 |
| 银屿 Yinyu Island | 5 | Vibrio alginolyticus、Vibrio owensii、Vibrio fortis | 3 |
| 全富岛 Quanfu Island | 6 | Vibrio alginolyticus、Vibrio owensii、Vibrio harveyi | 3 |
| 鸭公岛 Yagong Island | 9 | Vibrio alginolyticus、Vibrio harveyi、Vibrio mediterranei、Vibrio owensii、Vibrio variabilis | 5 |
| 甘泉岛 Ganquan Island | 7 | Vibrio alginolyticus、Vibrio coralliilyticus、Vibrio mediterranei、Vibrio variabilis | 4 |
| 羚羊礁 Antelope reef | 5 | Vibrio alginolyticus、Vibrio harveyi | 2 |
3 讨论
细菌种类鉴定是微生物领域的重要工作,近年来分子生物学技术的快速发展为微生物种类鉴定提供了技术支撑,分子生物学技术的稳定性、准确性可以很好地弥补生理生化鉴定的不足。16S rDNA序列分析法是细菌鉴定的基础方法之一,然而由于16S rDNA的高度保守性,使得16S rDNA序列在一些不同种细菌间的相似性非常高,甚至拥有完全一样的碱基序列[20]。Ohara C M等[21]和Erler R等[22]的研究表明16S rDNA序列不能区分弧菌属中的副溶血性弧菌和溶藻弧菌。本研究也发现,弧菌16S rDNA序列的种间相似度非常高,实验菌株与多个不同种模式菌株序列相似性同时高于99%,导致采用16S rDNA序列无法鉴定到种。
HSP60基因是高度保守的管家蛋白-分子伴侣蛋白编码基因,相比16S rDNA基因,HSP60的进化速度相对更快,因而更适用于作为近源关系细菌间鉴定的分子标记[23]。在本研究中也设计了一对HSP60引物,47株弧菌均能扩增出目标片段,但由于在GenBank中各类弧菌的HSP60基因信息量仍较少,相似度高于90%的弧菌仅有2种,因此目前仍无法利用HSP60基因准确鉴定弧菌到种。
gyrB为促旋酶的B亚单位基因,该基因进化速率快,每100万年的平均碱基替换率为0.7%~0.8%[24],属于信息通路中DNA复制、限制、修饰或修复有关的蛋白编码基因,存在于大多数细菌中,并且不显现频繁的水平转移,具备了作为系统发育分析靶基因的条件。由于gyrB作为蛋白编码基因,其所固有遗传密码子的兼并性使得DNA序列可以发生较多的变异而不改变氨基酸序列,因此gyrB基因在细菌的系统发育学,特别是在近缘种的菌株的区分及鉴定方面受到高度关注[25]。gyrB基因既弥补了16S rDNA的高保守性,也克服了16S-23S rDNA序列高变异性和复杂性。Venkateswaran K等[26]研究发现副溶血弧菌与溶藻弧菌的16S rDNA序列的相似率为99.7%,而1 258 bp的gyrB序列的相似率仅为86.8%,可以明显区分出这两种弧菌。Vuddhakul V等[27]通过对霍氏弧菌(V.hollisae)部分gyrB序列分析显示,该序列与副溶血弧菌的相似率为80%,可明显区分该两种弧菌。本研究选用16S rDNA、HSP60、gyrB作为靶基因对筛选的47株弧菌进行扩增及测序,通过与GenBank中相似性较高的弧菌进行同源性及系统发育树分析,结果发现16S rDNA和HSP60均不能成功鉴定弧菌到种,引物gyrB3能成功扩增所有47株菌株,且成功构建47株菌株的系统发育树,表明利用gyrB3引物进行海洋弧菌的分子鉴定是可靠有效的方法。
通过gyrB3的分子鉴定,在永乐群岛7个岛礁海域分离的47株弧菌分属于7个种,其中溶藻弧菌的存在具有普遍性,在7个岛礁海域中均有存在;而强壮弧菌仅在银屿海域中被分离到。在7个岛礁中,鸭公岛海域的弧菌多样性程度最高,甘泉岛、银屿、全富岛、晋卿岛次之,银屿和羚羊礁筐仔沙洲最低。近年来,永乐群岛的鸭公岛、银屿和全富岛开放了观光旅游,其中鸭公岛上还提供了游客的餐饮服务,餐饮垃圾与污水的排放可能是鸭公岛海域弧菌多样性较为丰富的主要原因之一。总体而言,永乐群岛7个岛礁近海海域的弧菌多样性较低。
参考文献
The depressed central carbon and energy metabolisms is associated to the acquisition of levofloxacin resistance in Vibrio alginolyticus
[J].
Metabolomics study of immune responses of New Zealand GreenshellTM mussels(Perna canaliculus) infected with pathogenic Vibrio sp.
[J].
Biodiversity of Vibrio
[J].Vibrios are ubiquitous and abundant in the aquatic environment. A high abundance of vibrios is also detected in tissues and/or organs of various marine algae and animals, e.g., abalones, bivalves, corals, fish, shrimp, sponges, squid, and zooplankton. Vibrios harbour a wealth of diverse genomes as revealed by different genomic techniques including amplified fragment length polymorphism, multilocus sequence typing, repetetive extragenic palindrome PCR, ribotyping, and whole-genome sequencing. The 74 species of this group are distributed among four different families, i.e., Enterovibrionaceae, Photobacteriaceae, Salinivibrionaceae, and Vibrionaceae. Two new genera, i.e., Enterovibrio norvegicus and Grimontia hollisae, and 20 novel species, i.e., Enterovibrio coralii, Photobacterium eurosenbergii, V. brasiliensis, V. chagasii, V. coralliillyticus, V. crassostreae, V. fortis, V. gallicus, V. hepatarius, V. hispanicus, V. kanaloaei, V. neonatus, V. neptunius, V. pomeroyi, V. pacinii, V. rotiferianus, V. superstes, V. tasmaniensis, V. ezurae, and V. xuii, have been described in the last few years. Comparative genome analyses have already revealed a variety of genomic events, including mutations, chromosomal rearrangements, loss of genes by decay or deletion, and gene acquisitions through duplication or horizontal transfer (e.g., in the acquisition of bacteriophages, pathogenicity islands, and super-integrons), that are probably important driving forces in the evolution and speciation of vibrios. Whole-genome sequencing and comparative genomics through the application of, e.g., microarrays will facilitate the investigation of the gene repertoire at the species level. Based on such new genomic information, the taxonomy and the species concept for vibrios will be reviewed in the next years.
一株大黄鱼致病性溶藻弧菌的分离鉴定与毒力相关基因分析
[J].2018年,宁德蕉城大湾渔排大黄鱼出现体表和鳍条溃疡、出血、鳃缺损等症状。为鉴定该菌株并评估其致病性,本研究从患病大黄鱼鳃部分离得到一株优势菌株,生理生化特征显示,该菌株可在TCBS培养基上生长,菌落颜色为黄色,为革兰阴性短杆菌,具有嗜盐性,生理生化指标与标准株17700一致。结合生理生化指标与16SrDNA 分析,确定该菌为溶藻弧菌,命名为Val180620。攻毒结果显示,Val180620菌株对大黄鱼具有较强的致病力,在水温25 ℃腹腔注射感染时,对规格(20±2)g大黄鱼的LD50为2.38×106 CFU。毒力相关基因分析结果表明,Val180620菌株携带有CollagenaseVA、ToxR、FlaA和TRH等毒力相关基因。此外,本研究未从菌株Val180620中检测出UreR基因,但菌株Val180620仍表现出较强的致病性,说明UreR不是决定溶藻弧菌毒力的主要因子。
创伤弧菌感染引起的坏死性筋膜炎2例
[J].创伤弧菌引起的坏死性筋膜炎具有高度的侵袭性和致命性,早期识别皮疹和干预可降低死亡率。本文报告创伤弧菌引起的坏死性筋膜炎2例。病例1,男,44岁,双下肢肿痛伴皮肤淤斑4天,既往有高血压病史和酗酒史。皮肤科查体:双下肢肿胀、广泛大片淤斑,部分张力性水疱、大疱和血疱。病例2,女,62岁,右上肢淤斑1天,腹痛和意识障碍半天。皮肤科查体:右上肢9 cm×10 cm大小的淤斑,上有个别水疱和血疱。2例患者的皮损分泌物培养均为创伤弧菌,结合临床症状,均诊断为创伤弧菌感染引起的坏死性筋膜炎。1例治疗无效死亡,1例放弃治疗,自动出院。提示四肢进行性淤斑和其上水疱或血疱可能是创伤弧菌引起的坏死性筋膜炎较早的危险信号。
Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence
[J].
Detection of bacterial 16S r RNA and identification of four clinically important bacteria by real-time PCR
[J].
The mcrA gene as an alternative to 16S rRNA in the phylogenetic analysis of methanogen populations in landfill
[J].
The all-species living tree project:A 16S rRNA-based phylogenetic tree of all sequenced type strains
[J].
中国明对虾急性肝胰腺坏死病(AHPND)和池塘水体中弧菌的检测分析
[J].近年来,对虾急性肝胰腺坏死病(AHPND)在全球范围内暴发,给对虾养殖业造成了严重的损失。为了进一步掌握中国明对虾AHPND的暴发与池塘水体中弧菌数量的关系,本文采用套式PCR法以及TCBS选择培养基法,分别对2018年4—9月采集的样品进行AHPND致病基因检测和池塘水体中弧菌数量的分析。套式PCR检测结果显示,7、8月AHPND检出率较高,分别达到15.6%、25.0%,其他月份未有阳性样品检出;TCBS选择培养基结果显示,在调水、控料等措施之下,4月至9月期间各虾池的弧菌数量控制在0~8.6×103 cfu?mL?1之内,其中7、8月达到高峰。另外,将本次试验中发现的TCBS黑色弧菌菌落,进行16S rRNA菌落PCR并测序,运用Mega 6软件中的Neighbor-Joining法构建系统进化树,结果显示其与溶藻弧菌(Vibrio alginolyticus)亲缘关系较近。本文针对该病的基本情况,结合近几年国内外的研究进展,提出了一些防控措施,为进一步控制该病害的发生提供了依据。
鳗源创伤弧菌的鉴定与血清型分析
[J].本研究对分离自发病鳗鲡的疑似创伤弧菌进行鉴定及血清型分析,经生化鉴定,从发病鳗鲡中得到7株创伤弧菌;设计了创伤弧菌溶血素基因(vlly)特异性引物,并对分离菌进行了PCR检测,创伤弧菌均可扩增出溶血素基因352 bp的目标片段,而非创伤弧菌未扩增出相应的片段;制备了创伤弧菌抗O抗原血清,凝集反应显示所得到的鳗源创伤弧菌抗O抗原血清不能与人源创伤弧菌1.1758发生凝集反应,菌株FJ03-X2 抗O抗原血清可与大部分的鳗源创伤弧菌发生凝集反应。以上研究结果表明,基于溶血素基因设计的PCR引物具有良好的特异性,可用于创伤弧菌检测,鳗源创伤弧菌血清型不同于其他来源的创伤弧菌,菌株FJ03-X2的O抗原具有良好的免疫原性,可作为创伤弧菌疫苗研发的候选菌株。
Ecological significance of microdiversity:identical 16S rRNA gene sequences can be found in bacteria with highly divergent genomes and ecophysiologies
[J].
Accuracy of six commercially available systems for identification of members of the family Vibrionaceae
[J].Six commercially available bacterial identification products were tested with Vibrio alginolyticus (12 strains), V. cholerae (30 strains), Photobacterium (Vibrio) damselae (10 strains), V. fluvialis (10 strains), V. furnissii (4 strains), V. hollisae (10 strains), V. metschnikovii (9 strains), V. mimicus (10 strains), V. parahaemolyticus (30 strains), and V. vulnificus (10 strains) to determine the accuracy of each system for identification. The products included API 20E, Crystal E/NF, MicroScan Neg ID2 and Rapid Neg ID3, and Vitek GNI+ and ID-GNB. Each product was tested only with those species that were listed in its database. Overall, the systems correctly identified 63.9, 80.9, 63.1, 73.6, 73.5, and 77.7% of the isolates to species level, respectively. Error rates ranged from 0.8% for the API 20E to 10.4% for the Rapid Neg ID3. The API 20E gave "no identification" for 13.1% of the isolates, while the Neg ID2, GNI+, ID-GNB, and Crystal were unable to identify 1.8, 2.9, 5.0, and 6.9%, respectively. For V. cholerae, specifically, accuracy ranged from 50.0 to 96.7%, with the API 20E having the worst performance and Crystal having the best. V. fluvialis presented the biggest challenge for the API 20E and the GNI+, with probabilities averaging 10%, while V. mimicus was a major problem with the Crystal E/NF, which identified none of the strains correctly. With the Neg ID2, correct answers were often obtained only after a modified inoculation of the panel with a bacterial suspension prepared with 0.85% NaCl. Additional tests required for identification often included growth in the absence of NaCl, which is not readily available in most clinical laboratories. The only product to correctly identify at least 90% of V. cholerae strains was the Crystal E/NF, and only three of the six products, the API 20E and both of the Vitek cards, correctly identified more than 90% of the V. parahaemolyticus strains. Thus, extreme care must be taken in the interpretation of answers from these six commercially available systems for the identification of Vibrio species.
Vibrio-Base:A MALDI-TOF MS database for fast identification of Vibrio spp.that are potentially pathogenic in humans
[J].
GyrB as a phylogenetic discriminator for members of the Bacillus anthracis-cereus-thuringiensis group
[J].
Cloning and nucleotide sequence of the gyrB gene of Vibrio parahaemolyticus and its application in detection of this pathogen in shrimp
[J].Because biochemical testing and 16S rRNA sequence analysis have proven inadequate for the differentiation of Vibrio parahaemolyticus from closely related species, we employed the gyrase B gene (gyrB) as a molecular diagnostic probe. The gyrB genes of V. parahaemolyticus and closely related Vibrio alginolyticus were cloned and sequenced. Oligonucleotide PCR primers were designed for the amplification of a 285-bp fragment from within gyrB specific for V. parahaemolyticus. These primers recognized 117 of 117 reference and wild-type V. parahaemolyticus strains, whereas amplification did not occur when 90 strains of 37 other Vibrio species or 60 strains representing 34 different nonvibrio species were tested. In 100-microliter PCR mixtures, the lower detection limits were 5 CFU for live cells and 4 pg for purified DNA. The possible application of gyrB primers for the routine identification of V. parahaemolyticus in food was examined. We developed and tested a procedure for the specific detection of the target organism in shrimp consisting of an 18-h preenrichment followed by PCR amplification of the 285-bp V. parahaemolyticus-specific fragment. This method enabled us to detect an initial inoculum of 1.5 CFU of V. parahaemolyticus cells per g of shrimp homogenate. By this approach, we were able to detect V. parahaemolyticus in all of 27 shrimp samples artificially inoculated with this bacterium. We present here a rapid, reliable, and sensitive protocol for the detection of V. parahaemolyticus in shrimp.
Analysis of gyrB and tox R gene sequences of Vibrio hollisae and development of gyrB and toxR targeted PCR methods for isolation of V.hollisae from the environment and its identification
[J].Isolation of Vibrio hollisae strains, particularly from the environment, is rare. This may be due, in part, to the difficulty encountered when using conventional biochemical tests to identify the microorganism. In this study, we evaluated whether two particular genes may be useful for the identification of V. hollisae. The two genes are presumed to be conserved among the bacterial species (gyrB) or among the species of the genus Vibrio (toxR). A portion of the gyrB sequence of V. hollisae was cloned by PCR using a set of degenerate primers. The sequence showed 80% identity with the corresponding Vibrio parahaemolyticus gyrB sequence. The toxR gene of V. hollisae was cloned utilizing a htpG gene probe derived from the V. parahaemolyticus htpG gene, which is known to be linked to the toxR gene in V. hollisae. The coding sequence of the cloned V. hollisae toxR gene had 59% identity with the V. parahaemolyticus toxR coding sequence. The results of DNA colony hybridization tests using the DNA probes derived from the two genes of V. hollisae indicated that these gene sequences could be utilized for differentiation of V. hollisae from other Vibrio species and from microorganisms found in marine fish. PCR methods targeting the two gene sequences were established. Both PCR methods were shown to specifically detect the respective target sequences of V. hollisae but not other organisms. A strain of V. hollisae added at a concentration of 1 to 10(2) CFU/ml to alkaline peptone water containing a seafood sample could be detected by a 4-h enrichment incubation in alkaline peptone water at 37 degrees C followed by quick DNA extraction with an extraction kit and 35-cycle PCR specific for the V. hollisae toxR gene. We conclude that screening of seafood samples by this 35-cycle, V. hollisae toxR-specific PCR, followed by isolation on a differential medium and identification by the above htpG- and toxR-targeted PCR methods, can be useful for isolation from the environment and identification of V. hollisae.